Let's Tackle the Nature H5N1 in Dairy Paper (Dimitrov-Diehl) in More Detail...
Broader herd coverage over time and geography adds clues
I missed by informal weekly deadline for a blog last week, electing to spend a weekend on the road with my 2 Ohio grandkids - that is never a bad decision!
Last week highly respected groups of researchers from Texas A and M, Cornell University, Ohio Animal Disease and Diagnostic Laboratory and Southeast Poultry Research Laboratory collaborated on publication of a review of spillover of highly pathogenic avian influenza H5N1 virus to dairy cattle in multiple herds in the U.S. in the early spring of 2024. The paper was published in Nature (link). Please read the paper in its entirety for much valuable case study information in multiple outbreak herds.
Data was collected on a total of 9 farms in multiple states, including one state (Ohio), where clinically normal cattle were shipped from a subsequently clinically infected Texas herd. I found the Ohio herd (#3) particularly interesting in that this was pretty clearly an imported incubating disease introduction, prior to any signs of clinical illness in the source herd. Because the cattle were shipped from Texas (herd #1) prior to initiation of clinical signs on March 9th (Extended Data Table 1) and the Ohio herd (#3) broke on March 21, that indicates a likely 11-14 day “incubation” period from arrival to onset of mastitis in the Ohio herd. The imported cows were lactating upon shipment, so apparently did not show sufficient mastitis or the herd did not collectively signal mastitis intervention until at least 11 days after arrival. According to the ARS mammary inoculation work (link), clinical signs of mastitis manifest approximately 48 hours post-inoculation. If the lactating cows shipped from Texas developed mastitis, shedding virus into the milking equipment, and infecting the Ohio herd, clinical mastitis was not noted in the imported cow herd for at least 11 days. Alternatively, the shipped cows may have carried a sub-clinical H5N1 respiratory infection, which was passed to the recipient herd over an 11-14 day period.
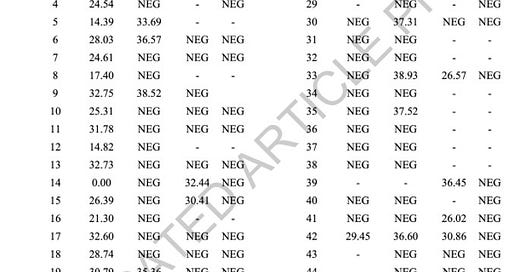
The paper actually makes a strong case for both clinical and subclinical respiratory and urine viral shedding by exposed animals. Here is a copy of Extended Data Table 2 from the paper, illustrating nasal swab and urine PCR titers were found in both clinically ill and subclinical animals tested in the study:
While PCR CT’s are not as strong as those found in milk, levels indicate definite viral replication with potential for intermittent shedding from large groups of both clinical and subclinical animals at level sufficient to allow transmission.
Here are comments from the authors:
Clinical animals shed virus at a lower frequency in nasal swabs (6/25) and urine (2/15), and no viral RNA was detected in feces. In non-clinical animals, viral RNA was detected in 6/19 nasal swabs and 4/8 urine samples, indicating subclinical infection.
Our results demonstrate a high tropism of HPAI H5N1 for the mammary gland tissue resulting in a viral-induced mastitis, which was confirmed by histological changes and direct viral detection in situ demonstrating viral replication and defining the virus tropism for milk-secreting mammary epithelial cells lining the alveoli in the mammary gland. The tropism of HPAI H5N1 for milk-secreting epithelial cells is consistent with high expression of sialic acid receptors with an α2,3 (avian-like receptor) and α2,6 (human-like receptor) galactose linkage in these cells36. Although the tissue sample size included in our study was small, isolation of the virus in lung and supra-mammary lymph nodes (which were also positive for viral RNA and antigen) suggests that other organs may also play a role in the virus infection dynamics and pathogenesis in dairy cattle. The initial site of virus replication remains unknown; however, it is possible that the virus infects through respiratory and/or oral routes replicating at low levels in the upper respiratory tract (e.g. nasal turbinate, trachea, and/or pharynx), from where it could disseminate to other organs via a short and low-level viremia. The collected evidence suggests that the mammary gland is the main site of virus replication, resulting in substantial virus shedding in milk. Another possible transmission route includes direct infection of the mammary gland through the teat orifice and cisternae, which could occur through contaminated floors and bedding where animals lay in the farm or mechanically via the milking equipment during milking. This entry route could also lead to viremia and subsequent virus dissemination/replication in other distant tissue sites. In the 1950’s several studies showed that direct inoculation of virus into the udder of dairy cows and goats with the human PR8 strain of type A influenza led to infection and viral shedding37–41. These results suggest, considering the current outbreak, that mammary epithelial cells, which express α2,3 and α2,6 sialic acid36, may be generally susceptible to influenza A viruses. There are a few studies suggesting an association between influenza A and clinical disease42–47, but there is no evidence of sustained transmission. The only published study of a goose/Guangdong lineage virus being inoculated intranasally into calves showed limited viral replication with no clinical disease and no evidence of transmissioin48. Experimental infection studies using different inoculation routes (i.e. intranasal vs intramammary) with HPAI H5N1 genotype B3.13 virus and, perhaps, other contemporary viruses of the 2.3.4.4b lineage in dairy cattle with sequential and comprehensive sampling are critical to answer these important questions about the port of entry, infection dynamics and pathogenesis in this new host species.
More complete commentary on the study featuring Dr’s Diego Diel (co-author) Richard Webby, and Carol Cardona can be found in the relatively recent New York Times article covering this paper: Halting the Bird Flu Outbreak in Cows May Require Thinking Beyond Milk - The New York Times (nytimes.com)
There is no doubt that the spectacular mastitis and extremely low H5 CT values in milk samples from this condition make the udder a very attractive target as the main site for understanding H5N1 in (dairy) cattle. It is full of the right receptors for triggering explosive replication of the virus. Milking cows also allow expression of loads of virus laden milk. However, making a big presence there (in perhaps 20% or less of an infected herd) does not mean the animal’s infection is initiated there. Injection through a teat orifice violates natural protective mechanisms normally at work. How many other herd-level systemic viral infections do we encounter with teat orifice introduction?
It’s this last step that still gives me pause… How do we, on a rapidly escalating basis, infect the udders of both lactating and nonlactating cattle (yes, astute vets have good clinical evidence of sick non-lactating cows, despite their inability to obtain serology testing), PLUS move the virus into shower-in high-biosecurity poultry facilities? The epidemic curve, variety of and sheer numbers of facilities infected, and the ongoing uptick in infected herds, despite all the efforts to tighten biosecurity, all lead me to conclude that udder infection and fomite spread just doesn’t fit the evidence for the outbreaks. An old-fashioned bovine respiratory influenza model with a 20% lactating cow mastitis sequella comes a lot closer to fitting the evidence as presented to date.
We all wish we didn’t have a new easily transmissible, potentially zoonotic cattle H5N1 influenza A virus. Trying to pretend it away won’t change the ultimate outcome. Let’s approve the needed serologic testing, conduct the robust transmission studies, and get on with testing the hypothesis and finding solutions!
John