OffFlu Group Proposals for Improved Livestock H5N1 Measures Will Likely Remain Underutilized Worldwide
Sparse mammalian livestock H5N1 surveillance is intentional in worldwide commercial agriculture with markets and business continuity to protect; white paper advocacy alone won't change that reality
Indiana Redux
Before launching into a separate topic, I’d like to provide a more recent update on the main topic in last week’s column. The Wednesday, March 25th, U.S. Animal Health Association Daily News Briefs featured the following:
5. Largest Animal Health Emergency in US History Continues as HPAI Hits Indiana Hard
By Eric Pfeiffer
WWBL.com
March 23, 2026
Since the beginning of March, highly pathogenic avian influenza has claimed over 350,000 birds in Indiana.
“We’re asking all of our producer partners, whether it’s layers or broilers or turkey or dairy or any other adjacent production system, please be vigilant,” says Dudley Hoskins, Under Secretary of Agriculture for Marketing and Regulatory Programs. “We need sound biosecurity practices. It’s not just what’s happening on that one facility, there’s risk of lateral transmissions.”
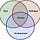
The Indiana tragedy was featured in my column last week; as of Thursday evening, March 26th, here is the updated March poultry count for Indiana:
The 350,000 birds listed about a week ago has now grown to nearly 400,000 depopulated poultry today.
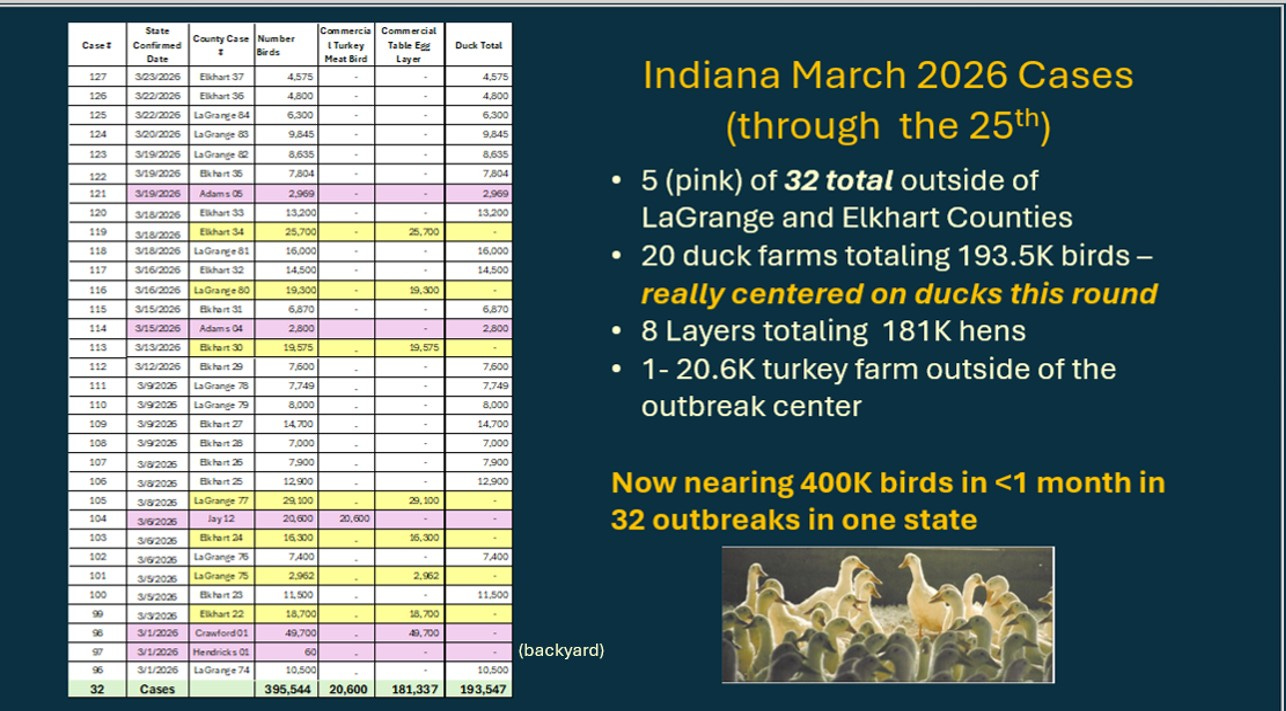
Here are the 2025-26 HPAI Indiana monthly totals by flock type:
March will go down as the second most deadly month of the 25-26 season in total poultry losses but the worst month for the duck industry. The state has lost over a million layers in multiple “smaller” flocks and over half a million ducks, predominantly in 2 counties in north Indiana! (3/31 update - Indiana BOAH has subsequently reported 4 more infected duck flocks totaling 27,120 birds through March 31st)
Moving briefly to another hard-hit state, I want to compliment Pennsylvania’s apparently quite successful effort to bring an ugly outbreak in Lancaster County under relatively rapid control! NVSL has only reported 3 Lancaster County PA outbreaks (plus one on neighboring Cecil County Maryland) since the 20th of March. I consider that really effective area outbreak shutdown, given the earlier scope of infection and the poultry density in that area. (3/31 update - NVSL reported 1 more infected Lancaster County duck flock totaling 18,000 birds on March 27th.)
Additionally, both Pennsylvania and Indiana have publicly stated that they are individually testing dairy herds (via milk sampling) in outbreak areas. No dairy herd infections have been reported in either state. It would be great if officials provided negative counts to document their dairy monitoring activities; however, the lack of reported cases is reassuring.
Mammalian Infections with H5N1 2.3.4.4b
State commitments to testing dairy herds in the immediate areas of poultry flock HPAI outbreaks remind us that this virus easily crosses into multiple mammalian species. I’ve included a couple of links to papers I reviewed this week that add to this body of information:
Detection of antibodies against influenza A viruses in cattle | Journal of Virology
This paper was published about a year ago but was not widely covered.
Unexpected outbreaks caused by the H5N1 highly pathogenic avian influenza virus (HPAIV) in dairy cows in the United States (US) have raised significant veterinary and public health concerns. When and how the H5N1 HPAIV was introduced into dairy cows and the broader epidemiology of influenza A virus (IAV) infections in cattle in the US remain unclear. Herein, we performed a retrospective study to screen more than 1,700 cattle serum samples collected from different bovine breeds in the US from January 2023 to May 2024 using an enzyme-linked immunosorbent assay (ELISA) targeting the nucleoprotein (NP) to detect IAV infections, and the positive samples were further tested by hemagglutination inhibition (HI) assay. Results showed that 586 of 1,724 samples (33.99%) from 15 US states were seropositive by the NP ELISA assay, including 78 samples collected in 2024 and 508 samples collected in 2023. Moreover, the HI assay revealed that 45 of these ELISA-positive samples were positive to human seasonal H1N1 and H3N2 and swine H3N2 and H1N2 viruses, and some were positive to two or three tested IAVs. Surprisingly, none of these ELISA-positive samples were HI positive for the circulating bovine H5N1 strain. Our results demonstrate that IAVs other than H5N1 can infect cattle, infections are not limited to dairy cows, and that bovine infections with swine and human IAVs have occurred prior to the H5N1 outbreaks. All results highlight the value in monitoring IAV epidemiology in cattle, as the viruses might adapt to cattle and/or reassort with the currently circulating H5N1 HPAIV, increasing risk to humans.
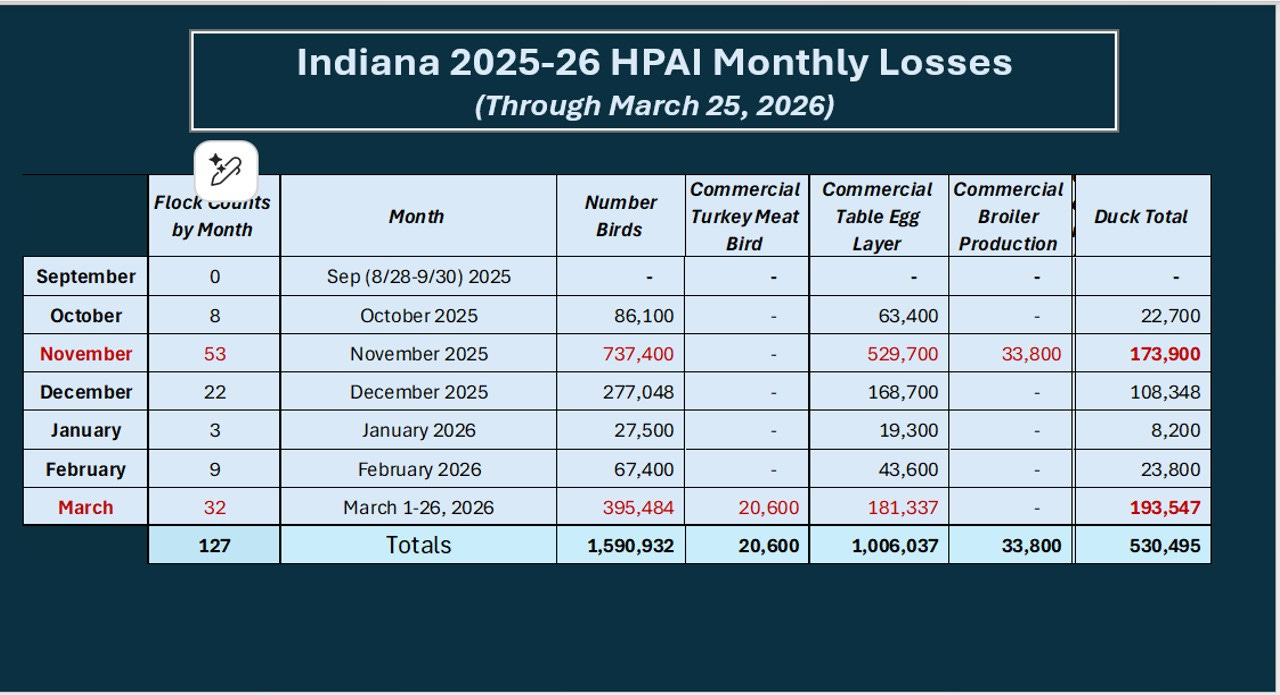
The results only reflect the time period sampled (January 2023-May 2024) and the areas from which the cattle were drawn (almost exclusively Missouri (1400+ samples), Illinois, Arkansas, Kansas, and Oklahoma-see map below). Lack of H5N1 titers is actually not surprising, given the May 2024 sample cut-off for a disease that first appeared in Texas in late March or early April 2024.
The more enlightening information from this paper is the high percentage of nucleoprotein (NP) ELISA positive samples, indicating influenza A exposure of some sort, with a significant number (45) showing HI titers to human and swine viral antigens.
The work provides further evidence of cattle susceptibility to influenza viral infection from multiple H subtypes and mammalian plus avian sources of virus. Further serological and antigenic studies, including H1, H3, and H5 across much broader populations of cattle would be useful, but unlikely to readily occur, as I’ll discuss later.
Michael Coston recently highlighted a report from Europe regarding detection of influenza A (H1N1) in a European bison: Avian Flu Diary: Vet Research Comms: First report of serological and molecular detection of influenza A (H1N1) in European bison (Bison bonasus). This adds one more piece to the body of evidence, including documented viral molecular detection, of the ability of influenza to at least sporadically infect ruminant species.
USDA Surveillance and Reporting of H5N1 Mammalian Infections
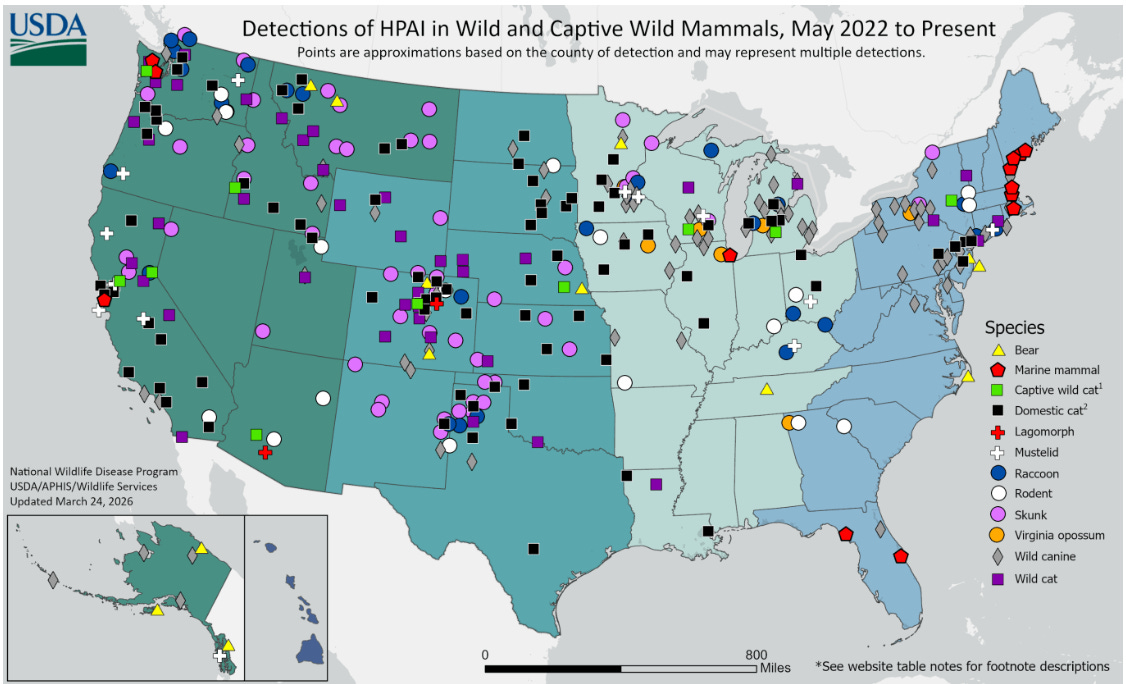
USDA has maintained a web site for some time now showing H5 isolates in “non-livestock” mammalian species in the U.S.: HPAI Detections in Mammals.
The dataset includes wildlife, including marine mammals, plus domestic cats. Sampling is based on passive and some active surveillance of affected animals, dependent upon voluntary mortality sampling conducted by federal, state, and local wildlife officials for the most part. Some samples may also be collected from peridomestic species collected near poultry and dairy HPAI outbreak sites. Positive results can reflect area incidence, but more likely reflects sampling intensity, or the lack of it, in many areas, e.g. most of the SE United States.
All positive farmed domestic mammalian livestock samples in the U.S. are found at this site: HPAI Confirmed Cases in Livestock | APHIS. With the exception of 1 backyard swine case in Oregon and 1 alpaca case in Idaho, all confirmed cases have been dairy cattle herds.
USDA-APHIS provides Testing Guidance for Influenza A in Livestock. Despite the growing scientific consensus for a wider sampling effort, this document is silent on testing production animals other than lactating dairy cattle. Slightly more guidance is found in the following on-line document:
Livestock HPAI TO (task orders) can be used for IAV-A testing in any species, including peri domestic mammals and birds, associated with a livestock premises. This includes cats. Any bovine IAV-A testing including FADIs can be charged to this TO.
What testing is paid for by NAHLN? Samples taken from other animals including cats with potential exposure (from dairies, exposed to raw milk, etc.) or reported clinical signs associated with this disease event.
Can NAHLN labs forward fluA non-negative samples from other livestock and companion species (e.g. horses) for characterization by sequencing? Yes. NVSL appreciates receiving these samples for further characterization.
Are positive test results for influenza A in all species reportable? All influenza A, with the exception of H1/H3 in swine should be reported to USDA.
In summary, USDA APHIS makes provisions for testing non-dairy livestock and cat samples for influenza through the HPAI “Dairy” Program at NAHLN labs and NVSL. However, those provisions are buried in a NAHLN “Dairy” FAQ document. Sample testing is not part of any coordinated approach towards risk-based sampling of livestock exposed to infected dairy herds, highly- infected HPAI poultry production localities, or to areas with high risks for wildlife H5N1 exposure.
OffFlu Discusses the Mammalian Livestock Surveillance Gap
Early last week OffFlu released a paper under the sponsorship of its parent organizations, WOAH and FAO:
The paper provides this opening Disclaimer:
This document does not provide normative standards or prescriptive guidance. It presents technical considerations and illustrative response options intended to support risk-based decision-making by competent authorities. This document provides the point of view of independent OFFLU experts and does not necessarily reflect the position of the parent organizations FAO and WOAH.
The introduction then is presented in summarized “Problem” and “Solution” summary statements:
Problem
Surveillance systems for high pathogenicity avian influenza (HPAI) urgently require adaptation to protect animal and human health in the context of a rapidly changing ecological risk landscape. This need is driven by the emergence and global spread of HPAI H5 clade 2.3.4.4b viruses of the Goose/Guangdong lineage (hereafter H5Nx), which have fundamentally altered the epidemiology of HPAI. H5Nx viruses have acquired increased fitness in wild aquatic birds, enabling sustained, year-round circulation and global spread. This ecological shift has led to recurrent incursions into poultry populations and a growing number of spillover events into diverse mammalian species, including companion and farmed mammals in close contact with humans.
The current surveillance paradigm, which remains largely focused on poultry, is no longer sufficient or reflective of the epidemiological reality. H5Nx viruses can no longer be regarded solely as “poultry-centered” or even “avian-centered”, highlighting the urgent need to expand surveillance efforts to include mammals. The continuous circulation of H5Nx viruses in wild aquatic birds for several decades suggests that these viruses are entrenched and unlikely to disappear in the near future. At the same time, it remains uncertain how ongoing viral evolution will affect host range, virulence, and transmissibility in different host species. This rapidly changing epidemiological landscape highlights the urgent need for versatile surveillance systems that are ecologically informed, cross species, and aligned with a One Health approach.
Solution
Many countries have well-established, regulated systems for the early detection and control of HPAI viruses in poultry, designed to prevent onward transmission and to minimize economic and animal health impacts. In addition, several countries have implemented surveillance programmes in wild aquatic birds to assess viral circulation, evaluate impacts on wildlife populations and ecosystems, and guide mitigation strategies aimed at reducing spillover into poultry.
However, dedicated HPAI surveillance strategies targeting wild and domestic mammals are largely absent in most countries. This gap is particularly critical for farmed mammals, which often live in close proximity to humans, are kept at high densities, are highly interconnected through trade and transport networks, and represent substantial economic value. Despite these risk factors, significant blind spots remain in monitoring programmes for HPAI virus infections in farmed (mammals), and in some settings such programmes are absent altogether.
A practical and scalable solution is to build upon existing avian surveillance systems that are already aligned with the WOAH Terrestrial Code and embedded within the One Health framework, by adapting these systems to include relevant mammalian hosts. ‘Relevant’ species are those whose biological susceptibility and ecological or geographical context place them at risk of involvement in H5Nx ecology, recognizing that what constitutes a ‘relevant’ species will continue to evolve over time and space due to viral evolution and changing exposure patterns.
Such integrated surveillance could aim both to detect H5Nx infections in mammals at an early stage and to prevent further spread, thereby reducing impacts on ecosystems, animal health, and human health.
The paper then provides more detailed background summary information, well worth a close read, before moving on to a section entitled “Detailed Solutions”, which the authors further break down across 4 broad groups: 1) mammals housed and managed in groups (e.g. livestock, fur farms, catteries, zoos), 2) individually managed companion animals, 3) free-ranging mammals (defined as limitedly- or non- managed mammals, living in free-ranging conditions, including both wild and feral mammals) and 4) humans. I’ll limit my thoughts to the group’s recommendations related to a subset of the first group- livestock and fur farms, where many of us are or were employed.
Please read the entire document closely - it is not excessively long. In zeroing in on the livestock-related recommendations, it occurred to me just what a heavy lift the group is requesting of the livestock industries in practice. Let’s look at some of the detailed recommendations to explore why few of these recommendations can occur without significant further international discussions and more creative surveillance paradigms.
Under the “Spillover” Prevention section the authors advocate for surveillance for “evidence of virus circulation in wild birds surrounding farms.” U.S. regulatory authorities have not provided for “real-time” reporting of H5 status in wildlife populations near farms; private monitoring for HPAI infection is not allowed. We can and do monitor wild bird population movements and densities and may get results of mortality testing for HPAI weeks or months after the fact. However, we lack assessments of the viral status of wildlife populations or the environment surrounding farms during periods of high risk. This is likely due to the risk that reporting a diagnosed wildlife HPAI infection in trade areas could adversely affect interstate or international trade in domestic poultry in some situations. H5 is now endemic to some degree in most areas in most countries involved in poultry trade; however, international agreements rules have not evolved to accept real-time wildlife status documentation for improved risk management without potentially penalizing reporting with possible trade restrictions!
Another statement in that section states: “Likewise, consideration may be given to reducing or phasing out intensive farming practices involving multiple species (e.g. poultry and pigs) on the same premises.” Multi-species single premise production may well increase risks for interspecies spillovers; however, recommendations for their phase-out would be viewed very dimly in the U.S. under our legal systems guarding free enterprise in agriculture. Our biggest multi-species on premises “violators” are likely our smaller producers! Additionally, repeated dairy to poultry premises to premises spread in 2024 demonstrated that understanding other factors in stemming “area spread” may be as critical as ending multi-species co-location on premises in mitigating cross-species spillover. Distance is not the only and perhaps not always the most important factor in limiting interspecies spread.
In the section titled “Early detection Through Targeted or Syndromic Surveillance”, the OffFlu group advocates for both broader testing of unexplained differential diagnostic cases, as well as targeted serological and PCR testing, especially in high-risk areas on appropriate slaughter animals and contact animals near where H5N1 is known to be circulating.
This section is where the good intentions and the most powerful recommendations of the OffFlu Working Group run head on into the realities of U.S. and international commercial agricultural production. Frankly, in the U.S. and EU, modern diagnostic methodologies and surveillance protocols and authorities are mature enough for full implementation of these recommendations. Next generation sequencing applied to “unexplained morbidity and mortality events (UMME) and targeted surveillance for HPAI in areas where HPAI is raging in poultry / wild bird populations have NOT been fully employed because we lack political will and compelling policies for implementation.
The paper claims routine implementation in the UK and the Netherlands; however, I have to wonder about its effectiveness in the Netherlands, given that nation’s track record on the H5N1 cat mortality-dairy farm traceback timeframe and associated dairy cattle PCR testing delays. I don’t bring this up to condemn the Dutch; I’ve also criticized U.S. officials for failing to test non-poultry species (swine and beef cattle) on quarantined HPAI premises. Zoonotic disease surveillance failures in livestock production units are a repeated occurrence in agriculturally mature developed countries. We can’t even pull raw cheese epidemiologically tied to E. coli infections (The Atlantic) from the market anymore!
We have little hope for significant consensus in sampling for H5N1 in livestock species (beyond our rapidly retreating dairy effort) until agricultural interests and regulatory officials are comfortable that sampling can be undertaken without jeopardizing interstate or international agricultural trade and market values.
The OffFlu paper moves on to suggest “Control of outbreaks through predefined response protocols”, stating “Suspicion (instead of detection) of H5Nx infection in any mammal could be made immediately reportable. The subsequent response measures could be determined based on risk assessment and local epidemiological context, rather than automatically prescribed.”
Key management options for consideration are similar to those implemented for other infectious diseases. Isolation may be applied, including defining minimum quarantine periods and clear criteria for release from isolation…
Culling may also be an option, and the benefits of preparing regulations for control measures in advance of outbreaks—rather than solely in response to outbreaks—may warrant consideration, with prioritization of the key species indicated earlier.
Ironically, the U.S. swine industry valiantly attempted to define many of these parameters for a possible H5N1 outbreak in swine herd(s) about a year ago, coming to consensus on an early draft for review by APHIS-VS. The effort basically went nowhere in APHIS, with so many questions outstanding regarding just how an uncharacterized novel agent of completely unknown zoonotic potential might need to be handled. My personal “take-home” was that it’s very difficult to prebuild a response plan for an H5N1 or reassorted virus of unknown infectivity and unclear zoonotic potential. Those factors alone make reporting infection or doing next generation sequencing of undiagnosed cases a tough decision, because the needed response is unknowable in advance of understanding the virus involved.
In the “Decrease impact when spillover has occurred” section, one final recommendation that will likely cause heartburn in the U.S. is: “Limits on animal numbers allowed on farms may improve control and response capacity…” Once again, economics and design solutions for large-scale viral outbreaks, whether HPAI or other diseases, will partly drive unit size decision-making. Ironically, ASF in China has seemed to drive even larger production units, right or wrong. Governments may choose to limit indemnification levels per site or emission levels per site; however, animal numbers per site based on disease risk is likely not the best way to solve an international diease problem.
The group made 4 final Considerations that work across these different phases:
• Acceleration of poultry vaccination efforts, when combined with appropriate biosecurity measures, could help to reduce overall viral burden and spillover risk.
• The development of best practices for mammalian influenza diagnostic techniques for viral (RNA) detection and antibody detection, including point-of-care diagnostics, could support improved detection capacity.
• Increased implementation of real-time genomic sequencing of viruses, alongside the sharing of genetic sequences from detections in all mammals (including free-ranging mammals) in the public domain, could support the rapid identification of adapted or reassortant viruses and enable genomic epidemiology, thereby improving risk assessment and response capacity.
• Consideration of inequalities and the establishment of financing mechanisms to support low-resource countries could strengthen the global response to this threat and support sustained reductions in overall viral burden and spillover risk.
I have supported all of these, especially 2 and 3; I’ve discussed both topics multiple times in multiple columns in the past.
An APHIS-coordinated NAHLN-based H5N1 integrated multi-species UMME and targeted surveillance process could be a powerful early detection and genotype monitoring tool in mammalian livestock species. It would meet most of the objectives of the OffFlu paper. I’m concerned that the current U.S. ‘Foreign Animal Disease Investigation” HPAI diagnostic monopoly is functionally limited in the production animal sector to poultry and dairy. It needlessly limits samples, ties up data, slows diagnostics, and prevents the widest and earliest possible dissemination of the broadest amount of H5NX 2.3.4.4b genomic information and associated metadata possible.
However, I also recognize that the agricultural meat and products trade industry may prefer to maintain the system just the way it is. We risk consumer or trading partner confidence in our meat and poultry products should unmonitored diagnostics in more species produce concerning results which are published “prematurely” from less controlled sources. Wide-spread cross-species H5N1 infection research studies and serological surveys are rarely conducted because approvals for conducting official tests in “off-target” livestock species must be granted by USDA-APHIS and resident state animal health officials. Neither individual states nor commodity group stakeholders want to volunteer as the next dairy industry H5N1 poster child!
Regardless, I personally believe that in the era of Oxford Nanopore and “Citizen Science”, trying to “manage” information through exclusive data channels is a longer-term futile proposition. We WILL have more H5 human hospitalizations/deaths and more sick livestock and wildlife as this virus keeps mutating and reassorting in birds and mammals. Viruses will be sequenced and associations made, some of them beyond the control of “official testing” performed exclusively in the regulatory-NAHLN-NVSL nexus.
Maybe it’s the optimist in me on behalf of my grandkids, but I’m increasingly confident that we are imminently approaching the onset of a resurgence in ethics and excellence in the U.S. and across the world that will sweep away much of the rot that currently governs us. Minneapolis showed the way for the rest of the world to follow. As this new mindset emerges across our society, we will collectively find the courage in agriculture to search out H5 and beat it, not bury it! Just like we beat brucellosis, TB, PRV, Salmonella enteriditis, and even E. coli in "no longer raw” cheese!
In U.S. animal health, we solve our problems, we don’t hide them!
John